And Holy Wars continues...
As a heroic adventurer of Blessed Python developed by our Noble Populace with Great Python Religion, I would like to mention few differences I noticed when working with Barbarous R. Few of my colleagues I regularly meet few Brutish Invaders of R, a land of primitive superstitions coming from Backward Savages.
This review is mostly about features regarding high performance mid-size (datasets about tenths of thousands rows you can fit in the 16GB memory) Data Science. And it is mostly for Data Scientists, not programmers (per se) or IT specialist. This article is also more a "brain dump" than something you would call a coherent report. And, of course, I write from a personal experience and not at all like an all purpose benchmark. I just wanted to summarized what I encounter, mainly for myself to remember.
By the way, I am not going to discuss feature parity. I think both languages/framework has big support of tools needed for ML/DS and this is not an issue today. Except neural networks - R's neuralnet is just a toy compared to Keras or Tensorflow. You really can't do much in this area with R.
Of course I may be wrong about many things in the following paragraphs - feel free to leave a comment and I promise will examine your objection.
Let's start
Both R and Python are available in Anaconda and that is positive. Nevertheless I used a compiled version of R from the official CRAN repository.
Coming from Python land, here is the list of stuff I found weird in R:
- No virtualenv by default - this was very limitting for me since I needed to benchmark different libraries with sometimes different versions. Later, I found packrat, which works but working with it is much less convenient than with
pyvenv - no multithreading by default and hard to actually do right - this strucked me most! When you use default CRAN version of R, you get it compiled with crippled BLAS implementation which doesn't support a multiprocessing even for embarrasingly paralelizable tasks, such as matrix multiplication. So everything is terribly slow. Forget about using more cores as is normal when you use numpy. GIL curse of CPython is nowhere close to this. If you wanna fast R, you need MRO or somehow link BLAS implementation when compiling R. Anaconda make this easier for you - you just need to download Intel MKL (which is great!). But I wasn't able to do this without root permissions on the test machine.
- readability worse (this is definitely a subjective and personal problem - deal with it)
- only
float64- so you can fit only half than withfloat32, or quarter in case offloat16or even eighth withint8that much into memory as you can with Python - overall worse memory management - in Python, I can fit 5e7 x 20 matrix into 8GB of RAM on dtype
float64(so as it should be:5e7 * 20 * 8 bytes = 8 GB). I can hardly do so in R - it seems it needs much more memory to allocate even random numbers. It can be garbage collected, but you need that memory to be able to generate that. This is huge. By allocating1e7 x 20matrix (so onfloat64:~1.7 GB), R allocates something like3.5 GB. So by factor 2. This can be tested by:matrix(rnorm(N*NCOL, ncol=NCOL),gc()(I was actually testing it withdata.frame(matrix...)). - generating random numbers take ages
Examples
Here are some examples how I measured the above:
Memory consumption
> N=2e7
> NCOL=20
> s = matrix(rnorm(N*NCOL), ncol=NCOL)
virtual/shared mem: 6198, 6137 (MB)
gc()
virtual/shared mem: 3146/3085 <- here we needed 100% more of mem
rm(s)
gc()
virtual/shared mem: 97KB/34KB <- correct, ~ 3.2GB
for 3e7:
> N=3e7
> NCOL=20
> s = matrix(rnorm(N*NCOL), ncol=NCOL) # takes 230 seconds
virtual/shared mem: 9.2/9.2 (GB)
gc()
virtual/shared mem: 4.6/4.6 <- here we needed 100% more of mem
rm(s)
gc()
virtual/shared mem: 97KB/34KB <- correct, ~ 3.2GB
but for Python:
# virtual/shared mem: 267MB,37KB (Ipython takes a lot, otherwise 97KB for vanila Python REPL)
import numpy as np
# virtual/shared mem: 357MB,52MB
s = np.random.randn(5e7, 20) # %time in Ipython
# virtual/shared mem: 8GB/7.7GB # <- almost no overhead, it took 62 seconds, for 3e7 it takes only 37 second
import gc
gc.collect() # doesn't change anything, is either not invoked or there is nothing to collect
del s
# virtual/shared mem: 430MB/53KB
so generating a Matrix of size 3e7 x 20 takes 6 times faster in Python and it needs only half the memory needed in R.
Logistic regression
init_data = data.frame(matrix(rnorm(N*NCOL), ncol=NCOL))
test_ratio = 0.7
test_split_n = round(N*test_ratio)
x_train = init_data[1:test_split_n, ]
x_test = init_data[-(1:test_split_n), ]
y_train = sample(c(0,1), length(x_train[,1]), replace=TRUE)
# 7.1GB/6.9GB
lr <- glm(y_train~., data=x_train)
the above exploded with RAM and swapped because it went over 16 GB, than garbage collected back to 7.1GB, than swapped to 29GB/13.4GB making the whole PC unusable and settled down on using about 2% of CPU (single core of course), while the status is Degraded. Stopped after 5 minutes of nothing (for scikit-learn 84 seconds with prediction). CTRL+C didn't work, had to kill it from the command line.
For 1e7:
# 3.6/3.6GB
lr <- glm(y_train~., data=x_train) # 69 seconds
# 6.8GB/6.7GB
e = predict(lr, x_test) # 3 seconds
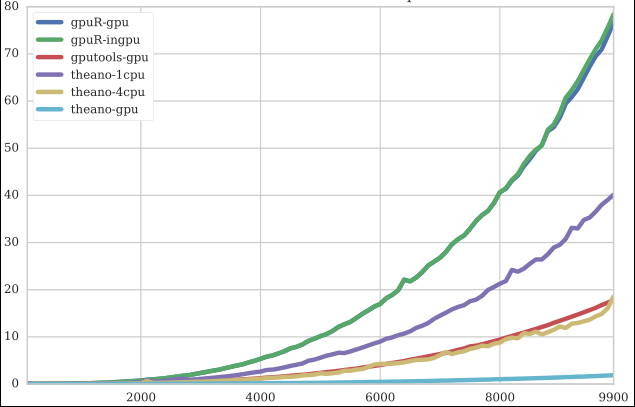
Comparison of GPU packages
I also looked over R's and Python's capabilities of GPU libraries. If you can read Czech, you may be interested in results in my research task (2016). Short version: R is slow. Even the fastest GPU implementation of various algorithms on R are slower than reasonable CPU versions in Python.
An example of matrix multiplication (CPU R version is 3 orders of magnitude slower, so not on chart):